 |
Web DSSR for dissecting the spatial structures of nucleic acids |
 |
| Home | Saturday, February 14, 2026 04:48 |
About
wDSSR (web DSSR) is a user-friendly web-based interface to the DSSR software package for the analysis and visualization of three-dimensional nucleic acid-containing structures, including their complexes with proteins and other ligands. Although this versatile and integrated suite of programs is widely used in the scientific community, its command-line-driven style is not especially user-friendly, for either novices (non-Linux/Unix users) or educational purposes.
Our new web-based interface provides straightforward access to some of the most popular features of the software, including:
Analysis:
The analysis component determines a wide range of conformational parameters — such as the identities and rigid-body parameters of interacting nucleic-acid bases and base-pair steps, the nucleotides comprising helical fragments, etc. — from a user-uploaded PDB-formatted file or a PDB ID. The output files can be viewed on the web or downloaded. Selected sequence and base-pairing information is reported in Grid-view tables with sorting and searching capabilities.
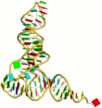
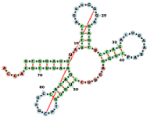
Visualization:
The visualization component creates secondary structure, allowing for simultaneously highlighting 1D, 2D, and 3D nucleic-acid structures. The server takes a user-uploaded PDB file or a PDB ID, and returns 1D, 2D, and 3D representations of the structure.
The server contains a database of pre-analyzed nucleic-acid-containing PDB structures to facilitate user access.
References:
3. Walther, D. (1997), WebMol: a Java-based PDB viewer. Trends Biochem. Sci., 22, 274-275.
4. JSmol: an open-source Java viewer for chemical structures in 3D. http://www.jmol.org.
Additional Readings:
Users interested in learning more about the content and capabilities of 3DNA should consult (i) the above references, (ii) the tutorial and worked-out examples found at the 3DNA website, and (iii) the user interchange at the 3DNA forum.
The following article describes the standard base reference frames, developed by the structural biology community and used in the 3DNA determination of rigid-body conformational parameters.
The conformational parameters for different types of nucleic-acid structures, reported in the following papers, provide useful benchmarks against which other structures can be compared.
The following papers pinpoint the differences among the programs most frequently used in the analysis of nucleic-acid structures: